 coelicolor to switch the superoxide dismutase (SOD) in response to oxidative stress under Ni-limited conditions. (B) Processing of the sodF mRNA by an unknown ribonuclease gives rise to the s-SodF sRNA. (A) The sRNA RaiZ derived from the 3′ end of the raiA mRNA acts in ProQ-dependent manner by suppressing the globally acting DNA-binding protein hupA affecting metabolism and virulence of Salmonella and E. By selectively inhibiting translation of nirC as part of the nirBDC-cysG operon, this sRNA works synergistically with the NarK protein to fine-tune nitrite uptake.įurther examples of 3′ UTR-derived sRNA in diverse organisms. (D) NarS is processed off the 3′ end of narK mRNA by RNase E. While SroC is involved in regulating motility by repressing translation of fliE, SroC has an expanded regulatory capacity through sponging the sRNAs GcvB and MgrR to further affect metabolism and LPS modification, respectively. (C)Premature transcriptional termination of the gltIJKL operon and subsequent RNase E-mediated cleavage frees the sRNA SroC from its parental operon. Additionally, SdhX exhibits divergent targetomes in E. (B) RNase E-dependent processing at the 3′ end of the sdhCDAB-sucABCD mRNA generates the sRNA SdhX, which selectively acts on ackA of the bicistronic ackA-ptA operon to regulate acetate metabolism. 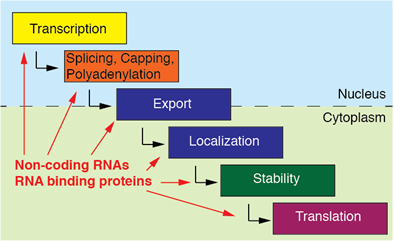
Additionally, in EHEC, DicF is also important for the regulation of the pathogen's virulence by upregulating the transcriptional activator PchA. Processing of the mRNA by both RNase E and RNase III yields DicF, which in turn is involved in the regulation of cell division and metabolism. coli ydfABC-dicF-dicB-ydfD operon mRNA that is transcribed from a defective prophage region. (A) The DicF sRNA stems from inside the E. Published by Oxford University Press on behalf of FEMS.Įnterobacterial 3′ UTR-derived sRNAs are involved in diverse pathways. Recent global RNA interactome studies suggest that the importance of functional 3' end-derived sRNAs has been vastly underestimated and that this type of cross-regulation between genes at the mRNA level is more pervasive in bacteria than currently appreciated.ģ′ UTR bacteria post-transcriptional control regulatory networks sRNA. Here, we review our current knowledge of these 3' end-derived sRNAs-their biogenesis through ribonucleases, their molecular mechanisms, their interactions with RNA-binding proteins such as Hfq or ProQ and their functional scope, which ranges from acting as specialized regulators of single metabolic genes to constituting entire noncoding arms in global stress responses. Several such 3' end-derived sRNAs have now been characterized, often revealing unexpected, conserved functions in diverse cellular processes. 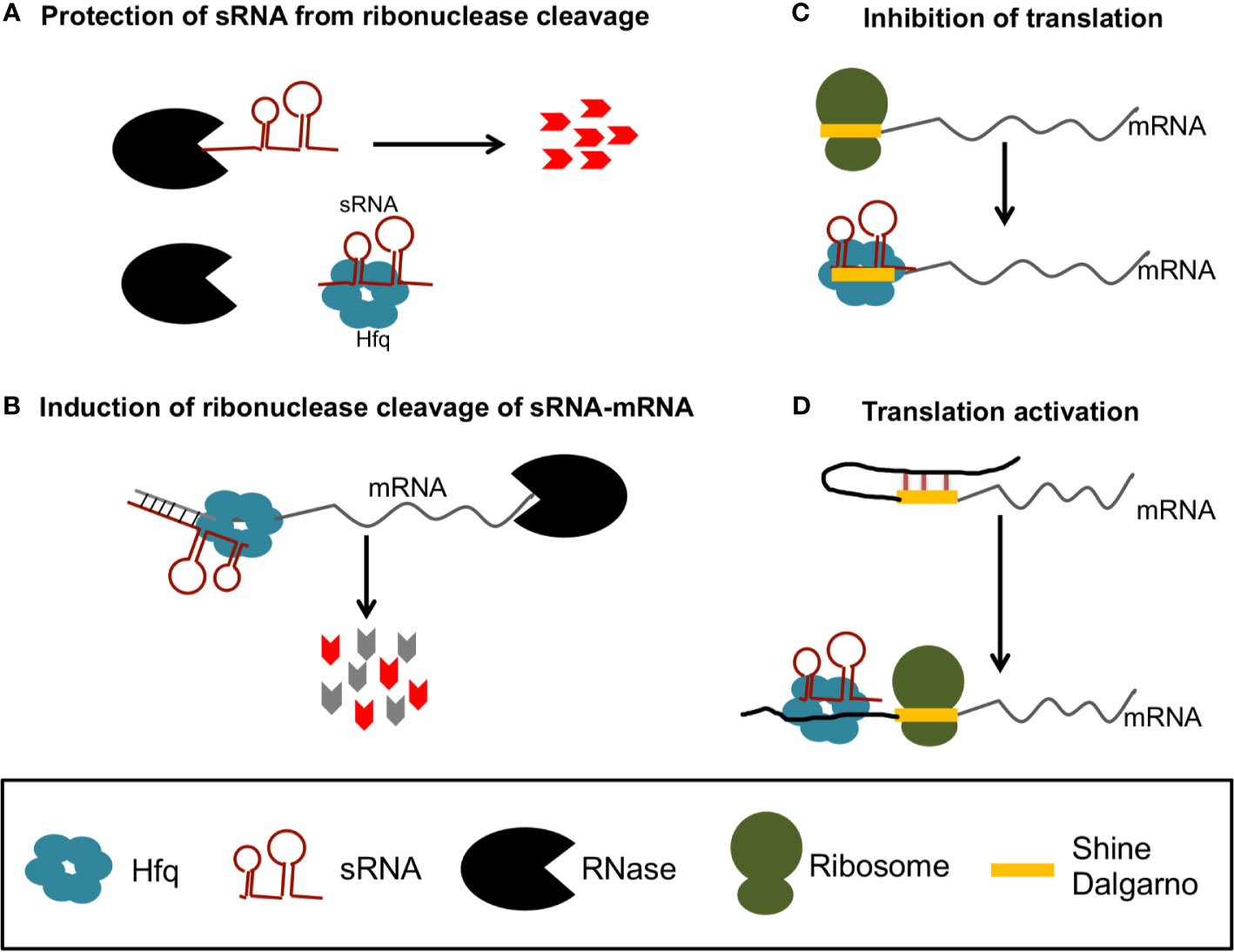
Yet, it soon emerged that there might be another rich source of bacterial sRNAs-processed 3' end fragments of mRNAs. Following pioneering sRNA searches in the early 2000s, the field quickly focused on conserved sRNA genes in the intergenic regions of bacterial chromosomes. 
They are integral to many stress responses and regulatory circuits, affecting almost all aspects of bacterial life. Over the past two decades, small noncoding RNAs (sRNAs) that regulate mRNAs by short base pairing have gone from a curiosity to a major class of post-transcriptional regulators in bacteria. 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
